The software, written in Python, enables automation of the analysis of epigenetic data from Illumina arrays that measure DNA modifications

Life Epigenetics, LLC – a leading developer of epigenetic technology for the longevity industry – has announced the release of two Python open source software packages to epigenetics researchers worldwide.
This software will facilitate scientific breakthroughs by accelerating and simplifying the processing of complex epigenetic data, the company said.
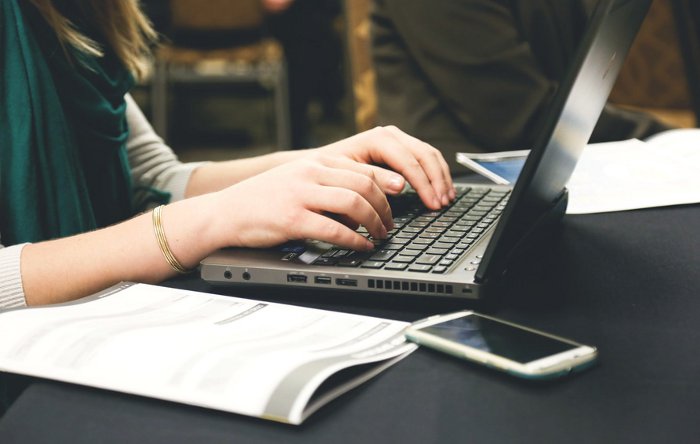
Essentially, the software enables automation of the analysis of epigenetic data from Illumina arrays that measure DNA modifications – specifically, adding or removing methyl groups – which are altered in response to human behaviour and biological processes.
With the software written in Python, the programming language most widely used by scientists and analysts, it can run natively in command line, Jupyter notebooks, or automation scripts.
These packages will provide improved user experience for the research community by automating previously laborious steps in epigenetic data pre-processing and quality control, the company said in a statement.
A gift to the research community
“We believe that by making this software available to any researcher, we will enable a whole new realm of collaboration and analysis at a critical time for epigenetics research,” said Life Epigenetics Lead Data Scientist, Dr. Randal Olson.
“Part of our mission is to openly share tools that accelerate research in epigenetics. By open sourcing this software, other researchers and developers can provide feedback and suggest ways to improve epigenetic data processing software for the worldwide research community,” Dr. Randal added.
These packages are the first of a series of open source software projects that Life Epigenetics plans to release over the coming year.
Through a series of partnerships and collaboration relationships, Life Epigenetics is working to commercialize groundbreaking epigenetic testing capabilities for insights into health and aging.