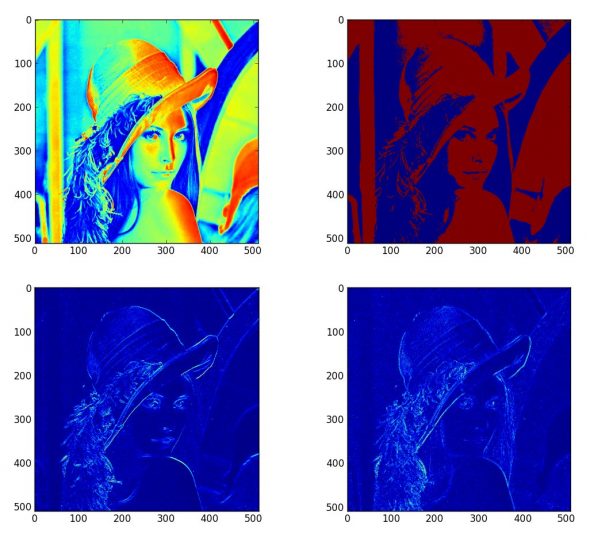
Discover what a beautiful language Python is for image processing.
A substantial part of the human brain is dedicated to vision and the processing of images. Social sites are full of images that friends share with each other. The sheer number of images from various sites is so huge that aggregation sites like Pixable hope to consolidate them for you, and help you find the ones you want from among all the clutter and noise. Another site, Scalado, created an application that allows you to remove unwanted people from your photographs, apart from various other things.
These examples illustrate that algorithms for the better classification or improvement of images and their content is a hot area. Python is a superb tool for exploring ideas and algorithms very quickly. While Python may seem unsuitable for computationally expensive tasks, the Scientific Python (SciPy) community has built tools for fast numerical computation while retaining the power, versatility and flexibility of Python.
A common option for image processing in Python has been the Python Imaging Library (PIL). However, implementing and exploring image-processing algorithms is better done in SciPy or NumPy. There are utility functions that convert a PIL image into a SciPy array and back:
im = Image.open('image.png')
A = scipy.array(im)
im2 = Image.fromarray(A)
But there is an ndimage module in SciPy, which makes it unnecessary to move between the two environments. The advantage of SciPy is that you can manipulate arrays in elegant ways, and not have to manipulate each element in a loop. Getting the sine of each element in an array, A, is as simple as what follows:
sinA = scipy.sin(A)
As you explore the possibilities with an image, you may find numerous uses for it in other areas as well, including financial modelling (read this, and this)!
Fancy indexing
The fancy indexing of NumPy/SciPy arrays provides a powerful abstraction tool. You can think about implementing algorithms at a higher level than manipulating elements of an array. The best approach is for you to actually try these in some simple but realistic examples, and judge for yourself.
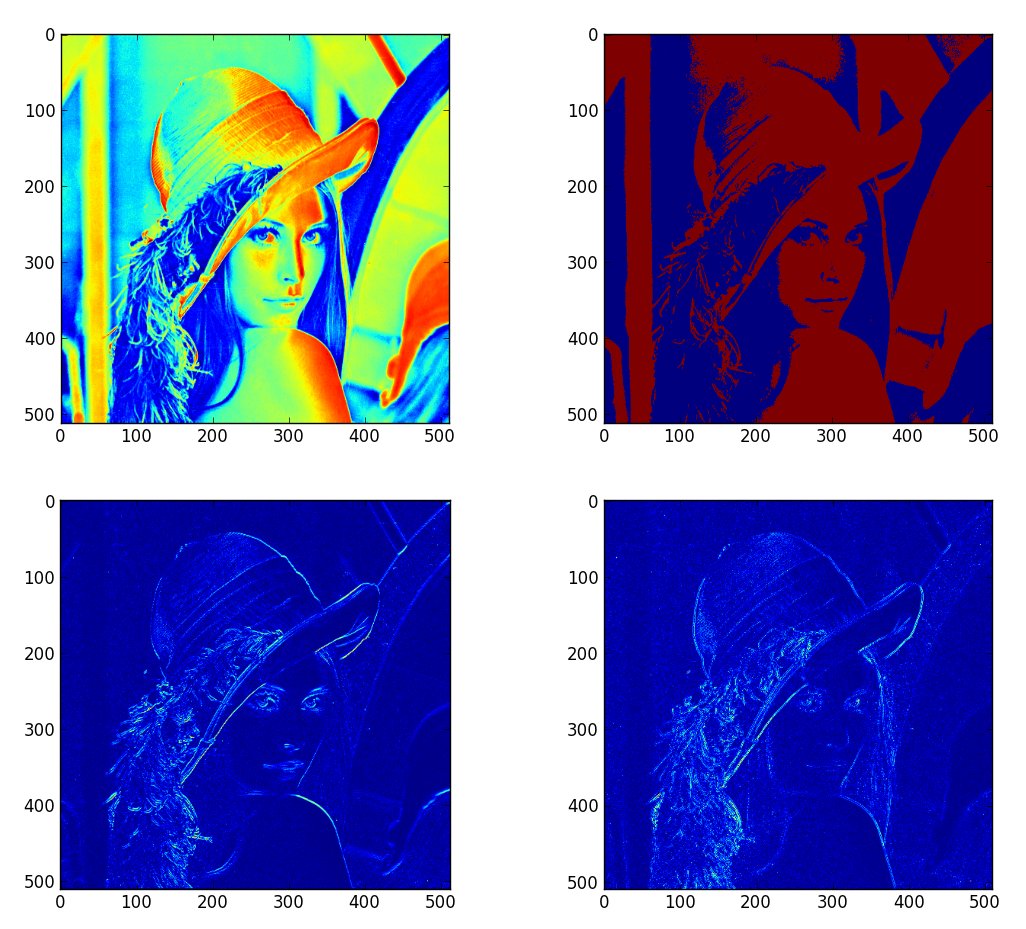
You may want to try implementing a threshold on an image, by using a reference image included in SciPy:
lena = scipy.lena() # create an array with zeros of the same size A = scipy.zeros(lena.shape, dtype='uint8') # find the elements in lena above the threshold, e.g. 120 mask = lena > 120 # set these to white (255) A[mask] = 255
The use of a mask as an index creates a very flexible and powerful way to manipulate arrays.
You could try a simple method to find a gradient in an image. One such relatively unused method would be to take the difference with the adjacent pixel in the same axis, like for a row. For example:
deltaA = A[1: , :] - A[:-1, :]
Indices are zero-relative, so the first term means a slice of the array starting with the second row, and all the columns. The second term represents a slice of the array starting with the 1st row till the last-but-one row, and all the columns. Both slices are of the same size, and can thus be subtracted. You get speed as well, because the element-wise operations are done by the C library.
Now, let’s assume that you wish to convolute an image with a 3×3 mask, M. This operation corresponds to element-by-element multiplication of the values in a 3×3 window around a pixel with corresponding mask elements, and then summing the result. For the sake of simplicity, you may ignore the border elements.
NR, NC = A.shape Result = [ [ (M*A[i-1:i+2, j-1:j+2]).sum() for j in range(1, NC-1)] for i in range(1, NR-1)]
You can use the Laplacian mask to get another image of the edges.
You will find a rich set of routines for filtering, morphology, segmentation and feature extraction in the ndimage module. To know more, just look at the section on image processing, analysis and face recognition (an example of machine learning).
New applications in image processing are waiting to be created. One wishes that Matlab would be replaced by SciPy in colleges — only then is it likely that a clever student will create a killer app.